 Consequently, devices with different degrees of automation have been developed. However, it can be time-consuming and suffer from a lack of reproducibility. 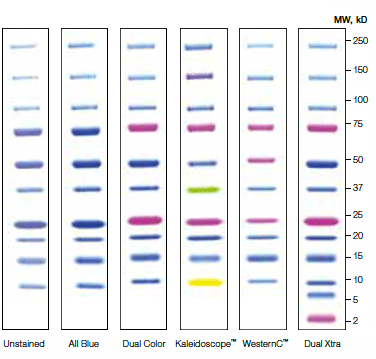
All the ladder proteins can be stained using dyes such as Coomassie Blue and are also directly detectable in Western blots through the incorporation of immunoglobulin antibody binding domains. Traditional Western blotting is one of the most used analytical techniques in biological research. This customization is easy to accomplish with the Penn State Protein Ladder system”įor even more efficient production, the ladders are also available as two coexpression vectors that produce either the set of 10, 30, 50, 100 kD proteins or the set of 20, 40, 60, 80 kD proteins. Accurate protein sizing of large molecular weight proteins on SDS-polyacrylamide gels and western blots. “They might not need the entire set, or they might want to individually control the intensity of bands produced on a gel. “Researchers can pick and choose amongst the nine individual proteins to design a ladder optimized for their research project,” said Tan. 50 milliliters (ml) of cells produce enough of each individual protein for 20,000 experiments. Proteins can be detected on Western blots by staining with Ponceau S, Coomassie Blue, (except for 3.4kDa and 5kDa polypeptides) or by using Strep- Tactin conjugates. coli cells containing the plasmids and then purify the proteins using a common affinity tag designed into each protein. Using standard laboratory protocols, researchers can grow E. Each protein is encoded on an individual plasmid-a circular form of DNA-that is expressed at very high levels in the bacteria E. The Penn State Protein Ladder is composed of nine proteins that range in molecular weight from 10 to 100 kilodaltons (kD). “Our Penn State Protein Ladders can be easily made for a tiny fraction of that cost using source material that we make available to nonprofit academic researchers through the Addgene and DNASU plasmid repositories.” “Any lab working with proteins uses these ladders on a daily basis and the cost for commercially available ladders adds up-averaging about $1.00 per experiment,” said Tan. Protein ladders are molecular rulers for estimating the sizes of proteins separated by gel electrophoresis. “It has been exciting and rewarding to develop this project from just an idea to tools that can serve science,” said Williamson. Fleischman, all of whom were Penn State undergraduate students. Williamson III, Yoshitaka Shibata, Rosalie P. Willaman Professor of Molecular Biology at Penn State, developed the ladders to be easily used in two of the most common experiments-SDS-PAGE and Western blots-in protein research.Ī paper describing the research appears Augin the journal Scientific Reports. Compatible with chemiluminescent substrates and fluorescent secondary antibodies.Labs can easily make their own protein ladders for less than a penny per experiment using the newly developed, license-free “Penn State Protein Ladder system.” A research team of undergraduate students led by Song Tan, Verne M. .jpg)
Someone recently told me that I should add Laemmli buffer to my ladder so that it is loaded at an equal.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
